Overview
Gene sequence processing involves collecting, reformatting, modelling and analysing complex and large sets of data (typically in 100's of millions) and presents technical challenges in choice of approach and technology. This page highlights example projects and technologies used primarily in vaccine design and analysis for HIV, Covid (SARS-CoV-2), Respiratory Syncytia Virus (RSV) and the Chikungunya Virus.
Processing gene sequence files (e.g. FASTA) often requires bespoke programs (often Python / Biopython) to reformat, align sequences, clean up and output required segments (e.g proteins or matching antigen regions).
Modelling using tools such as NetMHCpan can require parallel processing to maximise available resources and bring simulation times down to days rather than months.
Unix tools have proved fastest for post processing results as the low-level tools (e.g. grep, sed, awk) are suited for handling huge volumes of data. Formatting results into a SQL database allows for repeated analysis (excel is limited to a million rows). Python / Pandas is good for loading data into a database and any complex (than SQL reports) analysis scripts. Python / Biopython can also be useful in sequence analysis such as matching mutations and more complex ranking analysis methods.
Immunogenic Breadth Prediction Tool (IBPT)
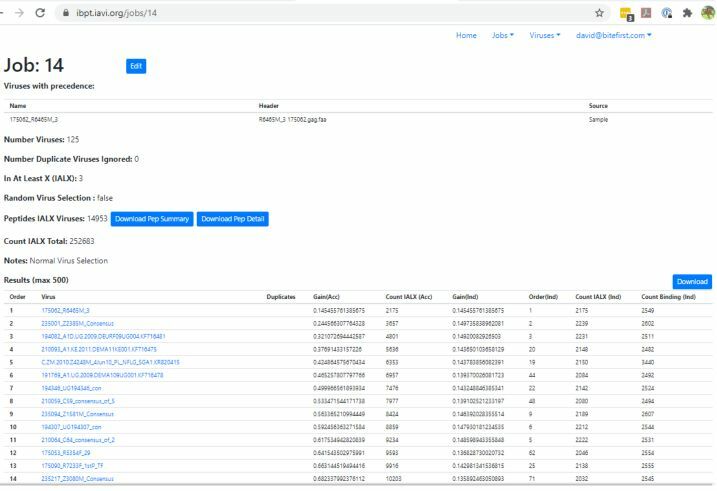
Example Analysis Run
The Immunogenic Breadth Prediction Tool (IBPT) was developed tool for IAVI. It is used to evaluate virus proteome diversity, as determined by prevalence of conserved Predicted CD8 T-cell epitopes, within IAVI Protocol C Transmitted Founder viruses and circulating HIV sequences from within LANL database (major USA database of HIV sequences).
The tool acts as a WEB based user interface to the results database (virus, hla, predicted epitopes and rank score) allowing for repeated analysis runs. e.g. help to identify region of a virus elicits a good T-cell response (most epitopes) and might be a candidate for a vaccine antigen.
The technology used is primarily Rails, Bootstrap, PostgreSQL with Python running on a nginx/thin services on an Ubuntu virtual server.
Example Results Set
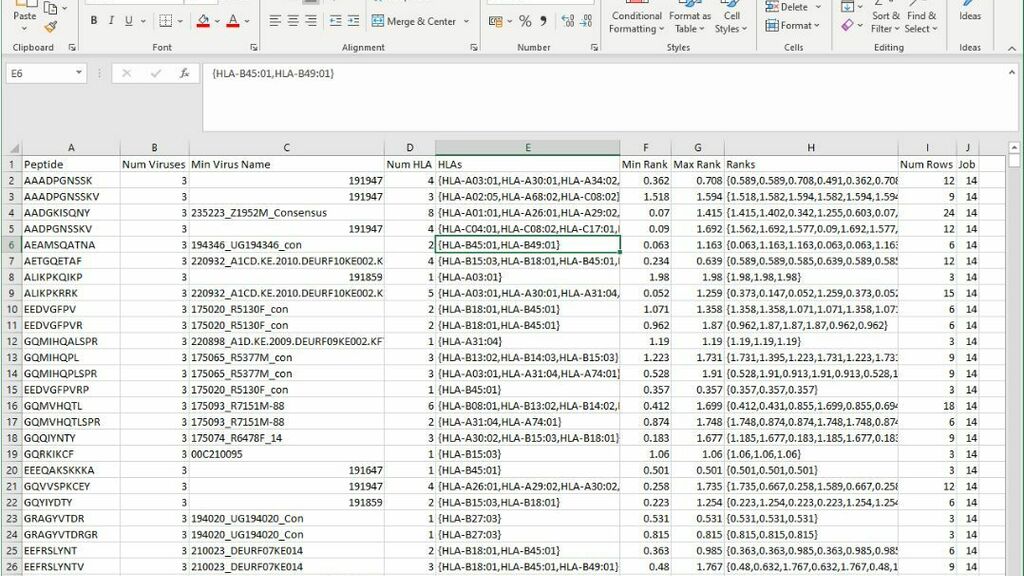
Example Summary
This is the output from a SQL query loaded into Excel that summarises the results. For a given predicted epitopes this shows the viruses that contain them, the HLA and rank score of the response. Large results files are stored on Google Cloud Storage (using Google Python API) which provides for large storage volumes and better handles download timeouts across poor network connections.
| Other Sections In This Site: |
|---|
| Sectors |
| Sectors / Clinical Data |
| Sectors / Bioinformatics |
| Sectors / Recruitment |
| Sectors / Billing |
| News |